
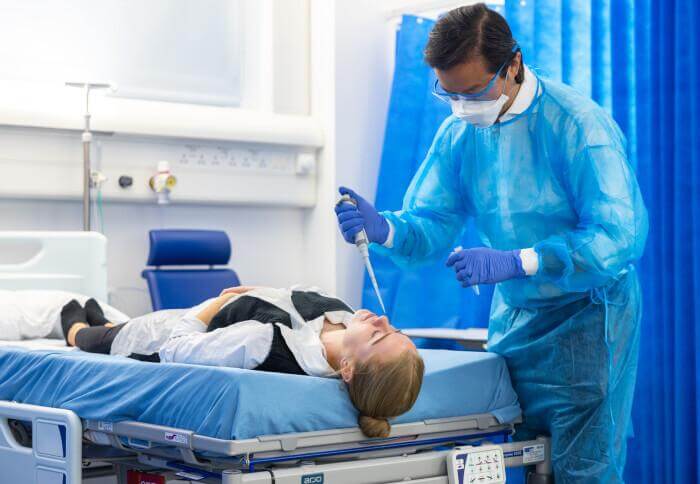
(© Dragana Gordic - stock.adobe.com)
CAMBRIDGE, United Kingdom — From mild cases to severe infections, hundreds of millions of people worldwide have had COVID-19 over the last four years. In the United States alone, the World Health Organization notes that one in three people have had a documented case of COVID. Despite the pandemic’s global reach, plenty of people have never gotten sick throughout this entire health crisis. Now, a new study may have finally discovered what keeps certain people from contracting this virus.
In an extraordinary leap forward in medicine, scientists have uncovered crucial immune responses that explain why some people exposed to SARS-CoV-2 manage to avoid developing the infection. This groundbreaking research, detailed in the journal Nature, used cutting-edge technology to map out immune dynamics in unprecedented detail, providing new insights that could lead to more effective prevention strategies.
Specifically, a team from the United Kingdom observed a never-seen-before immune response in people who immediately cleared the virus from their system without getting sick. Having higher levels of a gene called HLA-DQA2 before encountering SARS-CoV-2 protected these individuals from infections.
“These findings shed new light on the crucial early events that either allow the virus to take hold or rapidly clear it before symptoms develop. We now have a much greater understanding of the full range of immune responses, which could provide a basis for developing potential treatments and vaccines that mimic these natural protective responses,” says Dr. Marko Nikolić, senior author of the study from University College London, in a media release.
Methodology: Single-Cell Sequencing Leads to a Breakthrough
The study, conducted by researchers from the Wellcome Sanger Institute, University College London, Imperial College London, the Netherlands Cancer Institute, and other collaborating centers, employed single-cell sequencing — a method that allows scientists to analyze the genetic material of individual cells. This technique provided a detailed view of the immune system’s behavior in response to SARS-CoV-2 exposure by examining over 600,000 individual cells from the blood and nasal linings of 36 healthy adult volunteers. These volunteers, part of the UK COVID-19 Human Challenge study, were deliberately exposed to the virus, and their immune responses were carefully monitored afterward.
Results: Unseen Immune Patterns
The results of this study are nothing short of fascinating. Not all participants who were exposed to the virus went on to develop COVID-19, which allowed researchers to identify unique immune mechanisms that contribute to resisting the virus. Those who avoided sustained infection exhibited subtle, never-before-seen responses immediately upon exposure.
- Rapid immune response: The interferon response, a key early immune reaction, was noticeable in the patients’ blood before it was noticeable in the nasopharynx — the top of your throat connecting the nose and respiratory system. This suggests a pre-emptive systemic reaction that could be critical for controlling the virus.
- Diverse immune dynamics: Depending on whether the infection passed harmlessly or created a sustained illness, immune cells infiltrated the nasopharynx at different times, with a more immediate response in harmless infections, potentially pointing to a more effective containment of the virus.
- Role of specific genes: High expression of the HLA-DQA2 gene prior to encountering COVID displayed a link to preventing sustained infections, pointing to genetic factors that could influence individual outcomes to exposure.
Study Limitations
The controlled setting of the human challenge trial, while providing a unique opportunity to monitor the infection process from start to finish, does not entirely replicate natural exposure conditions. The findings are based on a relatively small sample size and a specific viral strain. Thus, while the insights are invaluable, further research is necessary to generalize these findings to broader populations and different SARS-CoV-2 variants.
Takeaways: A New Understanding of Immunity
This research maps the specific immune cells and responses that help some individuals spontaneously clear the virus without developing symptoms, paving the way for new strategies in vaccine development and pandemic preparedness. The discovery of HLA-DQA2’s role offers a potential biomarker for someone’s vulnerability to COVID-19, providing a pathway toward personalized medical treatments.
“As we’re building the Human Cell Atlas we can better identify which of our cells are critical for fighting infections and understand why different people respond to coronavirus in varied ways. Future studies can compare with our reference dataset to understand how a normal immune response to a new pathogen compares to a vaccine-induced immune response,” says Dr. Sarah Teichmann, senior author of the study and co-founder of the Human Cell Atlas, formerly at the Wellcome Sanger Institute, and now based at the Cambridge Stem Cell Institute.







