(© adimas - stock.adobe.com)
NIH’s SMaHT Network will chart trillions of somatic mutations to reveal how our DNA changes over a lifetime.
In a nutshell
- Scientists are creating the most comprehensive map ever of genetic changes that accumulate in your body throughout your lifetime—potentially quadrillions of mutations per person.
- Different body parts collect mutations at vastly different rates, with brain cells gaining 16-20 mutations yearly while colon cells rack up 44 mutations annually.
- This research could revolutionize medicine by helping doctors distinguish between normal aging changes and early disease signs, while revealing why identical twins become more genetically different over time.
BETHESDA, Md. — Your body is quietly collecting genetic changes right now, and scientists want to map every single one. A massive new research network is launching the most ambitious genetic study since the Human Genome Project, but this time they’re tracking the thousands of DNA alterations that pile up throughout your lifetime.
The Somatic Mosaicism across Human Tissues (SMaHT) Network, backed by the National Institutes of Health, plans to catalog genetic changes that happen after conception. These aren’t mutations inherited from your parents, rather, they’re DNA alterations that accumulate as you live, making each person’s tissues genetically distinct by adulthood.
These quadrillions of genetic changes help explain why identical twins grow more genetically different with age, and why diseases like cancer can seemingly appear from nowhere. “A typical cell may acquire hundreds to thousands of somatic mutations in a lifetime,” the researchers write in their paper published in Nature. “There are trillions of cells in a human body and so the total number of somatic mutations acquired in a single individual may well exceed quadrillions.”

Your Body’s Daily DNA Changes
Your cells face constant attack from environmental toxins, UV radiation, and normal cellular processes. Most DNA changes are harmless, but some dramatically alter cell behavior. Cancer represents the most extreme example, though these mutations also factor into heart disease, neurological disorders, and aging.
Different body parts collect mutations at wildly different rates. Brain cells, which rarely divide after birth, gather about 16-20 new mutations yearly. Rapidly dividing colon cells rack up approximately 44 mutations annually. Sun-exposed skin shows clear UV damage patterns, while smokers’ lung tissue bears molecular tobacco scars.
These patterns, called mutational signatures (distinct patterns that reveal what caused specific DNA changes), work like genetic fingerprints revealing specific environmental exposures or cellular processes that caused particular DNA changes.
“Specific mutations can be present in very small numbers of cells or even single cells, and so detecting them is like looking for a needle in a haystack,” says co-lead author Tim Coorens, from the Broad Institute of MIT and Harvard and currently a research group leader at the European Bioinformatics Institute, in a statement.
The Largest Genetic Study Ever Attempted
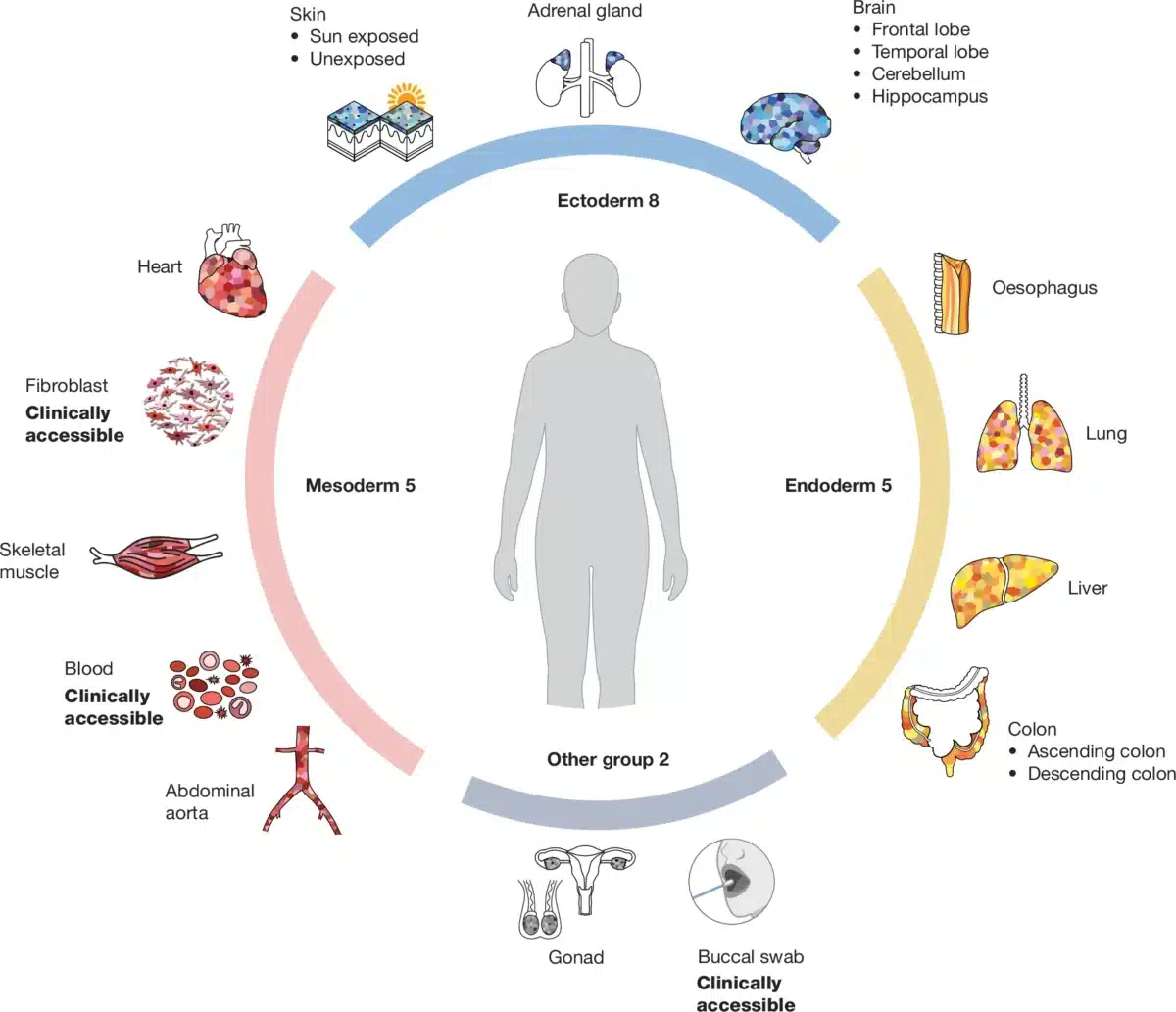
SMaHT researchers plan to analyze tissue samples from 150 deceased donors, examining 19 different tissue types from each person. Samples will include organs from all three developmental layers forming the human body: brain, skin, and adrenal glands; heart, blood, and muscle; plus lungs, liver, and intestines.
Advanced DNA sequencing methods will detect mutations present in just single cells among millions. One technique, duplex sequencing (a method that sequences both strands of DNA to reduce errors), reduces error rates to less than one in 100 million, allowing scientists to spot authentic mutations that would otherwise disappear in sequencing noise.
Over 250 researchers from 52 institutions are involved, organized into five genome characterization centers and 14 technology development projects. Each donor gets extensive analysis, with researchers collecting tissue samples plus detailed medical histories, environmental exposure data, and demographics.
Game-Changing Discoveries Already Emerging
Early SMaHT research has revealed unexpected insights about human development and disease. Studies tracking these mutations show that when a fertilized egg first divides, one of the two resulting cells often gives rise to twice as many descendant cells in the adult body as its sibling—explaining developmental patterns scientists observed for decades.
More surprisingly, normal tissues often harbor mutations typically associated with cancer. In healthy 60-year-olds, approximately 90% of endometrial tissue contains driver mutations that could theoretically cause cancer, while only about 1% of colon tissue shows similar changes, despite colon cells having much higher mutation frequency (the rate at which mutations occur).
This contradiction demonstrates how context determines whether mutations cause disease. The endometrium goes through monthly cycles of shedding and regrowth that may prevent dangerous cell populations from establishing, while colon tissue faces different biological pressures.
SMaHT’s comprehensive mapping could transform medical practice by establishing baselines for normal mutation patterns, helping doctors distinguish between harmless age-related changes and early disease signs. The research might also predict which patients face higher risks for specific cancers or other mutation-driven diseases.
Beyond medicine, the project promises insights into fundamental biological questions about aging, longevity, and how environmental exposures leave lasting cellular marks. Rather than having one static genome, each person carries millions of subtly different genetic variants distributed across tissues, a discovery that could revolutionize approaches to human health and personalized medicine.
Paper Summary
Methodology
Researchers from the SMaHT Network plan to collect tissue samples from 150 deceased donors aged 18 to over 85 years, examining 19 different tissue types representing all three developmental germ layers plus germ cells. They will use multiple DNA sequencing approaches including deep short-read whole-genome sequencing (300× coverage), long-read sequencing (30× coverage), and RNA sequencing. Advanced techniques like duplex sequencing will detect low-frequency mutations with error rates below 1 in 100 million. Single-cell DNA sequencing and transcript-based mutation detection will provide additional resolution. The project involves over 250 researchers from 52 institutions organized into five genome characterization centers, 14 technology development projects, and data analysis centers.
Results
Early SMaHT research has revealed that somatic mutation rates vary dramatically across tissues, from 16-20 mutations per year in neurons to 44 per year in colon stem cells. Different tissues show distinct mutational signatures reflecting their environmental exposures—UV damage in skin, tobacco signatures in lung tissue. Surprisingly, normal tissues often harbor cancer-associated driver mutations: 90% of endometrial tissue in 60-year-olds contains such mutations versus only 1% of colon tissue. Studies have also revealed developmental asymmetries, with one daughter cell from the first embryonic division often contributing twice as many descendant cells to the adult body.
Limitations
The study focuses on post-mortem tissue samples, which may not fully represent living tissue dynamics. Current short-read sequencing technologies limit detection of mutations in repetitive genome regions and may miss structural variations. Single-cell sequencing approaches can introduce amplification artifacts and uneven genome coverage. The project’s scope, while unprecedented, still represents a sampling of human genetic diversity and may not capture all possible mutation patterns across different populations or environmental exposures.
Funding and Disclosures
This research is supported by the NIH Common Fund. The authors report funding acknowledgments and any relevant financial disclosures in accordance with Nature’s policies. The study involves extensive collaboration between academic institutions and follows ethical standards required for human tissue research and donor consent.
Publication Information
This paper was published in Nature, Volume 643, pages 47-59, on July 3, 2025. The research was received on May 14, 2024, accepted on May 2, 2025, and published online on July 2, 2025. The work represents a collaborative effort by The Somatic Mosaicism across Human Tissues Network, with correspondence directed to Tim H. H. Coorens, Ji Won Oh, Eunjung Alice Lee, and Flora M. Vaccarino.







