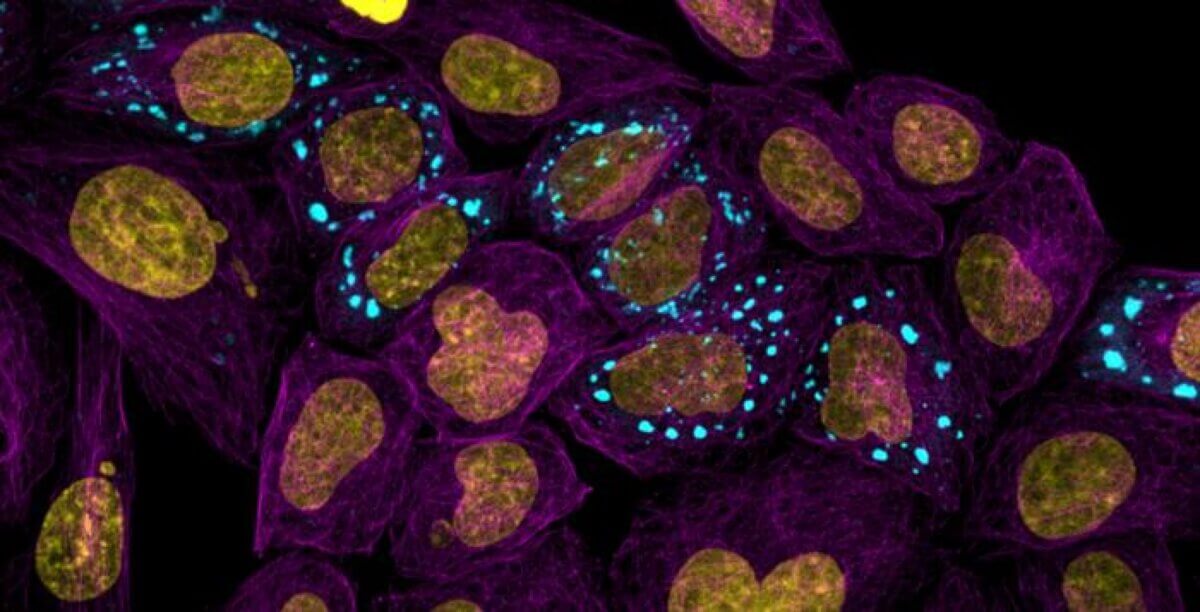
Microtubules highlighting the cell body (purple) with nucleus (yellow) and condensates (blue). (CREDIT: Andrew Seeber)
CAMBRIDGE, United Kingdom — Inside every living cell, a bustling metropolis of molecules is hard at work. For decades, scientists believed that the cell’s various components floated freely in a watery soup. However, recent discoveries have revealed a more organized picture – one where proteins and other molecules can spontaneously gather into droplet-like structures called biomolecular condensates. These tiny droplets act like cellular compartments without walls, allowing cells to organize their contents and carry out specific functions more efficiently.
Now, researchers from the University of Cambridge have developed a groundbreaking tool to predict and map these condensates across the entire human proteome – the complete set of proteins in human cells. Their work, published in Nature Communications, introduces the Protein Condensate Atlas, a comprehensive guide to the droplet-forming tendencies of human proteins.
“This model has allowed us to discover new components in membraneless compartments in biology as well as discover new principles underlying their function,” says Professor Tuomas Knowles, who led the research, in a statement.
Imagine a cell as a busy office building. In this analogy, biomolecular condensates are like impromptu meeting spaces that form when needed, bringing together the right proteins and molecules for specific tasks. These gatherings can regulate gene expression, help cells respond to stress, or assist in transmitting signals. Understanding which proteins tend to form these droplets and with whom could provide crucial insights into cellular functions and potentially lead to new therapeutic approaches for diseases linked to faulty condensate formation.
The research team set out to create a predictive model that could determine which proteins are likely to participate in condensate formation. They began by analyzing existing data on proteins known to gather in condensates, focusing on well-studied examples.
What they found challenged some previous assumptions about condensate formation. While certain protein characteristics, such as having disordered regions or low-complexity sequences, were thought to be crucial for condensate participation, the team discovered that these features were less important than expected. Instead, two key factors emerged as central to a protein’s tendency to join condensates: its electrical charge and its hydrophobicity (how much it repels water).
“What motivated this research was the desire to understand the full complexity of protein condensates and to go a layer deeper than what scientists have researched so far,” explains Dr. Kadi Liis Saar, first author of the research and a postdoctoral fellow at the Centre for Misfolding Diseases.
Armed with these insights, the researchers developed a machine learning model to predict whether a given protein is likely to participate in condensate formation. They then applied this model to the entire human proteome, creating a comprehensive map of potential condensate-forming proteins.
However, the team didn’t stop there. They realized that knowing which proteins tend to form droplets is only part of the picture. To truly understand cellular organization, they needed to predict which proteins are likely to gather in the same condensates. To accomplish this, they combined their condensate prediction model with existing data on protein-protein interactions, creating the Protein Condensate Atlas.
“To date we have not had a comprehensive map of which proteins go together into which condensates, but in our work we provide a first such atlas,” says Knowles.
This Atlas is like a social network map for cellular proteins, showing which proteins are likely to “hang out” together in droplet form. When the researchers compared their predictions to known condensates, they found striking similarities. Even more excitingly, the model predicted new proteins in condensates that had never been observed there before.
“With this atlas we can make predictions about every single protein in a cell, where exactly it would be found and what sorts of other proteins it interacts with,” explains Saar. “We hope that this generates opportunities for researchers and opens up new possibilities for intervention in diseases associated with aberrant condensate formation.”
The implications of this work are far-reaching. By providing a comprehensive view of protein condensate behavior, the Atlas could help researchers better understand how cells organize themselves and carry out complex functions. It could also guide the discovery of new drug targets, as many diseases, including dementia and various cancers, are linked to problems with condensate formation or regulation.
Moreover, the Atlas opens up exciting possibilities for exploring the uncharted territories of cellular organization. The researchers discovered proteins within condensates that have never been seen there before, involved in important functions such as fat distribution, actin creation inside cells, and the production of new proteins.
As our understanding of the cellular droplet universe expands, so too does our appreciation for the intricate and dynamic nature of life at its most fundamental level. The Protein Condensate Atlas represents a significant step forward in our ability to navigate and comprehend this microscopic world, promising new insights into cellular biology and potential breakthroughs in medicine.
Paper Summary
Methodology
The researchers used a multi-step approach to create the Protein Condensate Atlas. They analyzed existing data on proteins known to participate in condensates, identifying key features that distinguished these proteins from those that didn’t join condensates. They then used this information to train several machine learning models, testing different ways of representing protein sequences as input data.
The team utilized large databases like StringDB and BioGRID, which contain vast amounts of cellular data, along with more focused studies on individual condensates. The power of AI allowed them to integrate these complex and diverse datasets, enabling predictions about the entire cellular proteome rather than just a handful of proteins.
Key Results
The study found that protein charge and hydrophobicity were more important for condensate participation than previously thought factors like protein disorder. The machine learning model achieved good accuracy in predicting condensate participation.
When applied to the entire human proteome, the model identified many proteins likely to participate in condensates, some of which were previously unknown. The resulting Protein Condensate Atlas predicted numerous potential condensate clusters, many of which aligned well with known condensate systems. Importantly, the model discovered proteins within condensates that had never been observed there before, involved in crucial cellular functions.
Study Limitations
The study primarily used data from Western and White/European samples, limiting its generalizability to other populations. The predictions are based on existing protein interaction data, which may not capture all relevant interactions.
The model’s performance varied across different types of condensates, with some better represented than others. Additionally, while the model makes predictions for every protein in a cell, these predictions will need to be validated experimentally.
Discussion & Takeaways
This study represents a significant advance in our ability to predict, and map protein condensates across the human proteome. The Protein Condensate Atlas could serve as a valuable resource for researchers studying cellular organization and function. It may guide the discovery of new condensates and help identify additional components of known condensate systems.
The findings challenge some previous assumptions about condensate formation, highlighting the importance of charge-mediated and hydrophobic interactions. The study also demonstrates the power of combining machine learning with biological data to gain new insights into complex cellular processes. By making their predictions available to researchers worldwide, the team hopes to enable new discoveries about the biological roles of condensates and the biophysical drivers behind their formation. Future research could refine and expand the Atlas, potentially leading to new therapeutic approaches for diseases involving condensate dysfunction, such as dementia and certain cancers.







