Viruses know no borders. (Credit: Corona Borealis Studio/Shutterstock)
GRENOBLE, France — After nearly 40 years of research, scientists may be closer than ever to permanently knocking out the flu. Researchers in France have taken a groundbreaking look inside the intricate world of the influenza A virus, revealing how this microscopic invader packages and protects its genetic material.
For the first time, the study, published in the journal Nucleic Acids Research, offers an amazing view of how the virus constructs its internal structure, potentially opening new avenues for combating viral infections and making better medications for flu season.
Influenza viruses are notorious for causing seasonal epidemics and occasional pandemics, with the ongoing threat of avian flu strains potentially jumping from animals to humans. At the heart of the virus’s ability to spread and survive is its genetic material, carefully wrapped in a protein coat like a microscopic scroll.
Using cutting-edge imaging techniques, scientists have created the most detailed map yet of how the virus assembles its genetic material. The research focused on the nucleoprotein, a crucial protein that acts like a protective envelope for the virus’s RNA — its genetic instructions.
The team used a sophisticated approach to breaking down and rebuilding the virus’s protein structure in a laboratory setting. By strategically removing small portions of the protein’s structure, they were able to create more stable, less flexible versions of the virus’s core genetic package. This allowed them to capture incredibly sharp images that reveal how the genetic material winds through the protein’s structure.
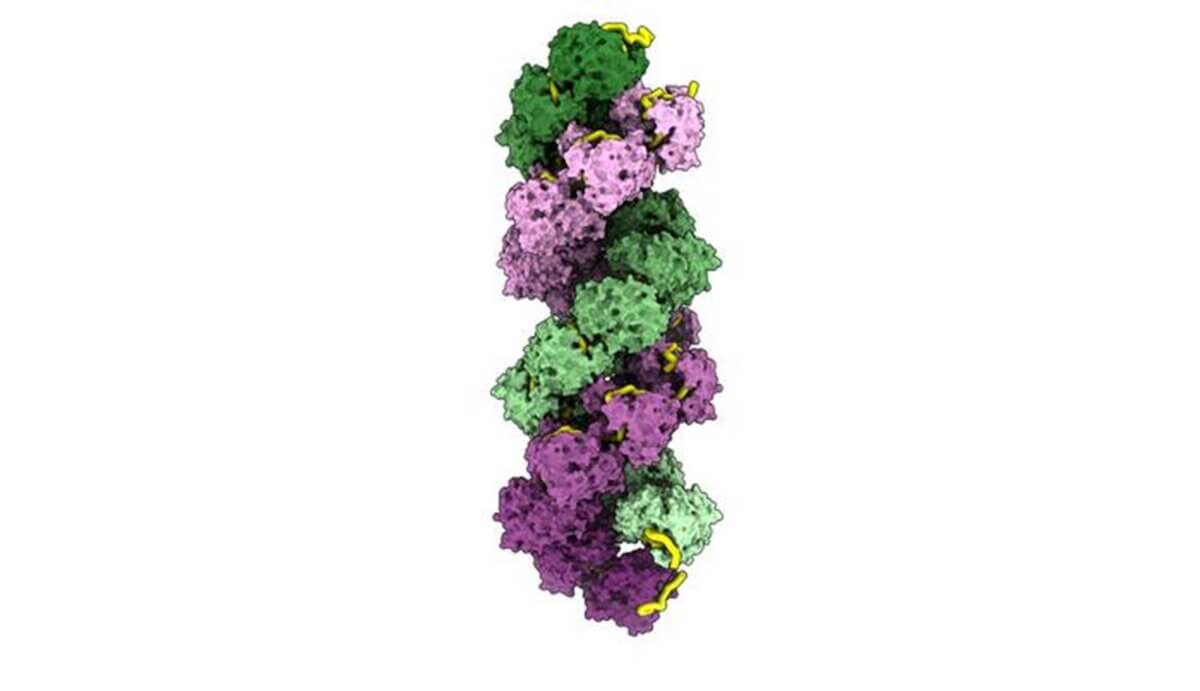
One of the most surprising discoveries was the flexibility of the virus’s genetic packaging. The researchers found that the protein can accommodate between 20 and 24 genetic building blocks, with one side of the protein more adaptable than the other. This flexibility might explain how the virus can quickly change and adapt — a key reason why influenza is so challenging to defend against each year.
The study also revealed intricate interactions between the protein and genetic material. The RNA (genetic material) doesn’t just sit passively inside the protein but actively helps shape the virus’s structure. It acts like both a cement and a spacer, contributing to the overall flexibility of the viral particle.
Importantly, the researchers noted that the protein’s first 14 amino acids play a crucial role in two key functions: providing flexibility to the virus and helping it move within host cells. This discovery could be a potential target for future antiviral strategies.
“This breakthrough paves the way for the design of new drug molecules capable of binding to the protein coat, weakening viral RNA, and inhibiting replication of the influenza virus, whose epidemics affect between 2 and 6 million people in France every winter and cause around 10,000 deaths in susceptible individuals,” the team from CNRS and l’Université Grenoble Alpes writes in a media release.
While the study was conducted in a controlled laboratory setting, it represents a critical step in unraveling the complex mechanisms of one of the world’s most adaptable and persistent viral threats. As influenza continues to pose challenges to global health, such molecular-level insights offer hope for more targeted and effective interventions.
For the general public, this research underscores the incredible complexity of viruses and the ongoing battle between human scientific innovation and microscopic pathogens. It’s a reminder of how much we’ve learned — and how much more there is to discover — about the tiny organisms that can have such a massive impact on human health.
Paper Summary
Methodology
In the research presented, the team developed a recombinant antiparallel helical nucleocapsid-like structure to explore the interactions within the influenza A virus’s ribonucleoprotein (RNP). This was achieved by using truncated forms of the viral nucleoprotein (NP) combined with synthetic RNA fragments. The core experimental technique involved cryo-electron microscopy (cryo-EM) to visualize the structures at a resolution of up to 3.0 Å. Different constructs of the NP were engineered, expressed, and purified, allowing the researchers to observe how variations in the NP contributed to the formation and stability of the nucleocapsid-like structures.
Key Results
The study successfully mapped the intricate details of the influenza A virus’s RNP architecture by employing cryo-EM, revealing that partial deletion of the NP N-terminal tail stabilizes the nucleocapsid-like assembly. This manipulation facilitated a detailed view of NP-NP and NP-RNA interactions, highlighting the protein’s flexibility and its implications for RNA encapsidation. Specifically, the researchers found that varying the length of RNA affected the formation of single-stranded and double-stranded helices, with certain lengths fostering more stable interactions and assembly formations.
Study Limitations
One limitation of the study is the use of truncated versions of the NP and synthetic RNA, which might not fully replicate the natural interactions and conformations present in the viral RNP of influenza A. Furthermore, while cryo-EM provides significant structural insights, the resolution limitations mean that some interactions at the atomic level might not be fully discerned. The extrapolation of these findings to understand the full virus’s behavior in biological systems thus requires careful consideration and further validation.
Discussion & Takeaways
The findings underscore the flexibility of the influenza A virus’s nucleoprotein as crucial for its ability to encapsidate RNA effectively, a factor that might influence viral replication and host adaptation. The ability to modify the NP to stabilize different helical forms provides a valuable tool for further studies on virus assembly and pathogenicity. Understanding these mechanisms could potentially lead to novel antiviral strategies that target the structural components of the virus.
Funding & Disclosure
The research was supported by the Chemistry Biology Health Graduate School of University Grenoble Alpes and the Agence Nationale pour la Recherche. The authors declared no conflicts of interest, ensuring the study’s impartiality and integrity in presenting unbiased findings.







