Credit: NMK-Studio on Shutterstock
Scientists identify measurable cellular changes that flag higher cancer risk more than a decade before tumors appear.
In A Nutshell
Liver cells under chronic dietary stress essentially “forget” their normal jobs, shutting down essential functions to focus on bare survival. It’s a trade-off that creates cancer-prone conditions years before tumors appear.
The warning signs show up early: Molecular changes in liver cells can flag elevated cancer risk 10-15 years before hepatocellular carcinoma develops, potentially opening a window for early intervention.
One enzyme tells the story: HMGCS2 levels decline during metabolic stress, and lower levels in human liver tissue predict both disease progression and future cancer risk.
Future screening may look different: Instead of waiting for tumors on scans, doctors might eventually use molecular signatures from biopsies to identify high-risk patients when cellular dysfunction first begins.
Liver cells overwhelmed by dietary fat essentially forget how to be liver cells, according to research. Under chronic stress, they progressively shut down the genes that define their normal responsibilities and switch into a bare-bones survival mode. And these changes, the study shows, can signal elevated cancer risk years before any tumor appears.
Scientists at MIT and Harvard tracked this cellular transformation in real time, watching liver cells reprogram themselves over 15 months as mice consumed high-fat diets. The findings, published in Cell, reveal that the body’s short-term strategy for keeping liver cells alive under constant dietary assault creates conditions that make cancer far more likely down the road.
The discovery could reshape how doctors monitor the more than 33% of people globally who have metabolic liver disease. Instead of waiting for tumors to show up on scans, physicians might eventually use molecular fingerprints from liver biopsies to identify high-risk patients when cellular dysfunction first begins—potentially a decade or more before cancer develops.
The Cellular Trade-Off Nobody Wants
Consider a liver cell as an employee at a company who makes critical products for the entire organization. Under normal conditions, that employee handles hundreds of specialized tasks, processing nutrients, manufacturing proteins the body needs, cleaning up toxins.
But under relentless stress from excess dietary fat, that same employee faces an impossible choice: keep doing the specialized work that benefits the whole organization, or focus entirely on personal survival. The cells choose survival.
Stressed liver cells ramped down production of the proteins and enzymes that perform the liver’s signature jobs. They made less of the enzyme controlling ketogenesis (the process of converting fat into fuel for other organs) and the urea cycle (which handles nitrogen waste). They cut back on albumin and clotting factors, proteins the blood carries throughout the body.
At the same time, they activated an entirely different playbook. They switched on gene programs that resemble early liver development and cranked up proteins that block cell death. They increased certain cholesterol-making enzymes while decreasing the ketone-producing ones, even though both pathways work from the same raw materials.
After 15 months on high-fat diets, some mice spontaneously grew liver tumors without any genetic manipulation or cancer-causing chemicals. Those tumors showed even more extreme versions of the same cellular reprogramming.
Warning Signs That Appear First
Incredibly, the cancer-associated changes appeared long before actual tumors.
At just six months of dietary stress, liver cells already showed signs of preparing the ground for future problems. Specific regions of their DNA (ones that control genes involved in cell growth and cancer) became more accessible, like files that had been pulled from storage and placed on a desk, ready to be opened. These regions stayed poised for months until tumors eventually formed.
When researchers examined human liver tissue from patients at different stages of fatty liver disease, they found the same progression. People with early-stage disease already showed activation of genetic programs characteristic of liver tumors. More tellingly, the strength of these early signatures was linked to which patients developed hepatocellular carcinoma, the most common liver cancer, up to 15 years later.
The same gene programs appeared in liver cancers that arose from different causes: metabolic disease, viral hepatitis, and alcohol-related damage, suggesting common pathways through which diverse types of chronic injury may contribute to cancer formation.

One Enzyme Connects the Dots
Of all the changes the team documented, one enzyme stood out: HMGCS2. This protein normally runs the first critical step in ketogenesis, helping the liver convert fat breakdown products into ketone bodies that fuel the brain and muscles when food is scarce.
HMGCS2 levels dropped steadily as mice stayed on high-fat diets. When scientists created mice genetically engineered to lack this enzyme in liver cells, those animals showed dramatically accelerated cellular dysfunction. More critically, they were far more vulnerable to tumor formation when exposed to cancer-causing genetic changes.
In human patients, lower HMGCS2 in non-cancerous liver tissue linked to both worsening liver disease and higher risk of eventual cancer. The enzyme’s decline appears to be both a result of chronic stress and an accelerant of further problems. Without enough ketogenesis happening, metabolic intermediates may pile up and get shunted into processes that alter how genes are read and expressed, potentially helping explain how dietary stress rewires cellular behavior.
Molecular Switches That Tip the Balance
To figure out what controls this widespread cellular reprogramming, researchers built a computational tool that predicted which molecular master switches might be calling the shots.
Two proteins that control gene activity, SOX4 and RELB, emerged as key players. Normally quiet in adult liver, both became more active as metabolic disease worsened in mice and humans.
When scientists artificially boosted SOX4 in liver cells, it triggered many of the same changes seen during chronic dietary stress: cells activated fetal development programs, suppressed mature liver functions, and kept dividing even under conditions that would normally stop proliferation. Higher SOX4 and RELB in non-cancerous liver tissue was associated with worse outcomes in patients who eventually developed cancer.
From Research to Real-World Application
The findings point toward a fundamentally different approach to cancer prevention in high-risk patients. Rather than relying on imaging that spots tumors after they form, doctors might one day measure a panel of molecular markers –HMGCS2 levels, SOX4 and RELB activity, and specific gene program scores — to stratify patients by cancer risk.
Some existing treatments may already affect these pathways. Resmetirom, recently approved for metabolic liver disease with scarring, targets a molecular switch that the computational analysis flagged as important for these stress responses.
Of course, there is one big unanswered question. Can these cellular changes be reversed? Weight loss and newer medications like GLP-1 receptor agonists improve liver tissue appearance, but researchers don’t yet know if they erase the deeper molecular reprogramming that occurred during months or years of metabolic stress. The elevated cancer risk might linger even after the liver looks healthier.
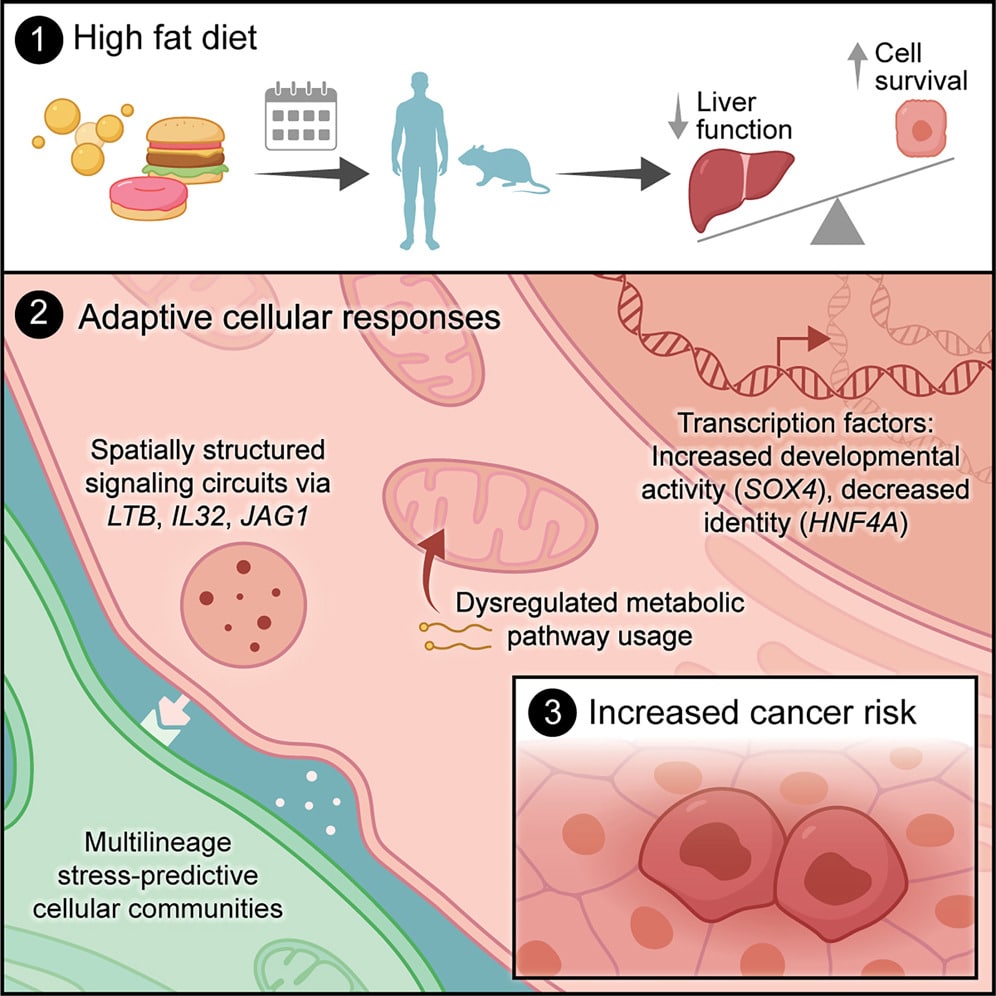
The Bigger Picture
When cells face prolonged stress, they can’t do everything at once. They make trade-offs between their specialized roles and their own survival. Those trade-offs work in the short term: they keep cells alive when they might otherwise die. But maintained over months and years, they leave behind a liver that’s functional enough to sustain life but riddled with cells that are primed for cancer.
The fact that these early adaptations leave measurable traces opens up the possibility of catching the very first steps toward cancer, years before the first tumor cell appears, when intervention might still change the outcome.
Disclaimer: This article describes research findings about cellular changes in metabolic liver disease and their association with cancer risk. It is intended for informational purposes only and does not constitute medical advice, diagnosis, or treatment recommendations. Individuals with fatty liver disease, metabolic concerns, or questions about liver health should consult qualified healthcare providers for personalized medical guidance and appropriate monitoring options. The research described involves laboratory studies and observational human data; specific screening tests or interventions mentioned are not yet widely available in clinical practice.
Paper Notes
Limitations
The mouse model used constitutive liver-specific knockout of HMGCS2 beginning during embryonic development, which could have led to compensatory developmental changes. The model also did not develop significant bridging fibrosis comparable to human disease, possibly due to differences in mouse versus human lifespan. Additional limitations include that the high-fat diet model likely involves multiple physiological changes beyond direct nutrient overload, including potential effects from insulin resistance, gut-liver crosstalk, adipose-liver signaling, and other systemic factors. The spatial transcriptomics analysis was conducted on cirrhotic liver tissue, representing late-stage disease. Questions remain about the timescales of stress response initiation, persistence, and potential reversibility with interventions.
Funding and Disclosures
C.N.T. is supported by a Fannie and John Hertz Foundation fellowship and a National Science Foundation Graduate Research Fellowship. J.E.S.S. is supported by a National Institutes of Health F32 Fellowship, NIH grant P30 DK040561, and the AGA Research Foundation’s Research Scholar Award. M.S.S. is supported by NIH grants T32DK007191 and K08DK139370. W.G. is supported by NIH grants R01DK090311, R01DK105198, and R24OD017870. Ö.H.Y. is supported by NIH grant R01CA245314. A.K.S. is supported by NIH grant DP1DA053731. Multiple authors receive support from the MIT Stem Cell Initiative through Fondation MIT. A.K.S. reports compensation for consulting and/or SAB membership from multiple biotechnology companies unrelated to this work. C.N.T., J.E.S.S., M.S.S., W.G., Ö.H.Y., and A.K.S. have filed a patent related to this work.
Publication Details
Authors: Constantine N. Tzouanas, Jessica E.S. Shay, Marc S. Sherman, and multiple co-authors from Massachusetts Institute of Technology, Harvard Medical School, Brigham and Women’s Hospital, Dana-Farber Cancer Institute, Ragon Institute of MGH, MIT and Harvard, Koch Institute for Integrative Cancer Research at MIT, and Broad Institute of MIT and Harvard.
Journal: Cell | Article Title: Hepatic adaptation to chronic metabolic stress primes tumorigenesis | Published online December 22, 2025 | DOI: 10.1016/j.cell.2025.11.031







