
Credit: sergey kolesnikov on Shutterstock
Study Shows Drug Factories Are Pumping Superbug Genes Into Rivers, And Nobody’s Stopping Them
In A Nutshell
- Antibiotic factories pump out concentrated resistance pollution. Manufacturing plants show resistance genes in waste at levels 100 times higher than city wastewater treatment facilities, creating ideal conditions for superbugs to evolve and spread.
- The pollution spreads through rivers and soil into food systems. Factory waste contaminates downstream waterways and agricultural areas, embedding resistance genes in environments where they can persist and transfer to bacteria that cause human infections.
- Technology exists to eliminate 90% of resistance genes from factory waste. High-heat composting, biological treatment systems, and targeted gene-destruction technologies are proven to work, but no regulations require pharmaceutical companies to use them.
- Pharmaceutical facilities face almost no monitoring for biological pollution. While hospitals and farms are tracked by global surveillance programs, antibiotic factories lack standardized protocols for measuring resistance genes in their environmental discharge.
Investigations into antibiotic-resistant bacteria contaminating rivers and soil have revealed an overlooked pollution source. The factories that manufacture antibiotics themselves.
Drug manufacturing plants pump out wastewater packed with antibiotic residues and bacteria that have learned to survive those drugs. Per the research, published in Biocontaminant, one plant in China showed resistance genes measured in its activated sludge at levels 100 times higher than municipal wastewater treatment plant sludge.
Scientists warn that those resistance genes can spread to bacteria that cause human infections. The result is superbugs that shrug off the antibiotics doctors prescribe. Between 2025 and 2050, antibiotic resistance could directly kill 39 million people.
The Pollution Nobody Talks About
Most people know antibiotics come from pharmacies. Fewer realize they come from massive industrial facilities that produce antibiotics at global scale.
The manufacturing process creates a problem. When antibiotics are produced industrially, it generates enormous volumes of contaminated water. That water contains leftover drug ingredients, chemical waste, and most dangerously, bacteria capable of surviving in an environment saturated with antibiotics.
An aforementioned plant in China that produces spiramycin showed resistance genes at concentrations over 100 times higher than municipal wastewater treatment plant sludge. Even worse, those genes had attached themselves to what scientists call “mobile genetic elements,” which are essentially biological USB drives that let bacteria copy and share resistance instructions with each other.
Where Antibiotics Factory Waste Goes
Researchers studying a sulfonamide drug plant collected samples from the river downstream and surrounding areas. They found that the pollution had traveled. Soils and river sediments showed high levels of resistance genes. These measurements exceeded global averages.
The contaminated soil posed a particular concern. The pollution wasn’t washing through, it was embedding itself in the environment.
The agricultural connection matters because rivers water crops and soil contamination affects food-growing regions. The genes that help bacteria survive antibiotics can potentially move from factory waste into agricultural systems.
Scientists found high-risk resistance genes and bacteria resistant to multiple drugs throughout these downstream areas.
Why Antibiotics Factory Pollution Is Different
Of course, not every source of antibiotic pollution is equal. Large-scale antibiotic production creates waste streams with high concentrations of antibiotics and resistant bacteria. This creates intense selection pressure, an evolutionary environment where only the toughest bacteria survive.
Bacteria that survive pharmaceutical manufacturing conditions carry genetic tools to defeat multiple antibiotics simultaneously. When these supercharged strains escape into rivers or contaminate soil, they can teach other bacteria their tricks. That includes bacteria that cause human infections.
The river doesn’t care whether bacteria came from a hospital patient or a factory drain. Once they’re in the environment, they’re all mixing together, swapping resistance genes.
The Missing Regulations
Hospitals and farms face monitoring for antibiotic resistance through global health programs. Additionally, The World Health Organization’s surveillance system and agricultural monitoring platforms track resistance patterns in clinics and livestock operations across countries.
However, pharmaceutical manufacturing facilities, which can contain some of the highest concentrations of resistance genes, lack standardized protocols for monitoring their environmental impacts. Current regulations measure chemical pollution, like leftover drug ingredients. They rarely address the living pollution: bacteria and resistance genes that can reproduce and spread.
Scientists lack comprehensive global data on pharmaceutical facilities’ contributions to environmental resistance, despite these being among the most concentrated sources.
What Could Be Done
Solutions exist, but aren’t being utilized.
Special high-heat composting processes can eliminate about 90% of antibiotic resistance genes from pharmaceutical waste. Specialized bacteria can break down antibiotic residues before they create selection pressure for resistance. Some newer technologies can even target and destroy specific resistance genes.
None of these solutions are theoretical. They work in research settings. They’re just not required.
Why This Matters to You
If you get a routine infection (a urinary tract infection, strep throat, a skin infection from a cut, etc) doctors prescribe antibiotics expecting them to work. As resistance spreads, those routine infections become harder to treat. Some become untreatable.
The resistance genes flowing out of pharmaceutical plants can eventually end up in bacteria that cause common infections. Rivers carry contamination downstream. Soil contamination affects agricultural systems. The pollution pathways run through water systems and into food supplies.
This isn’t about far-off threats in distant countries. Pharmaceutical manufacturing operates as a global supply chain, and pollution from these facilities can contribute to resistance problems worldwide.
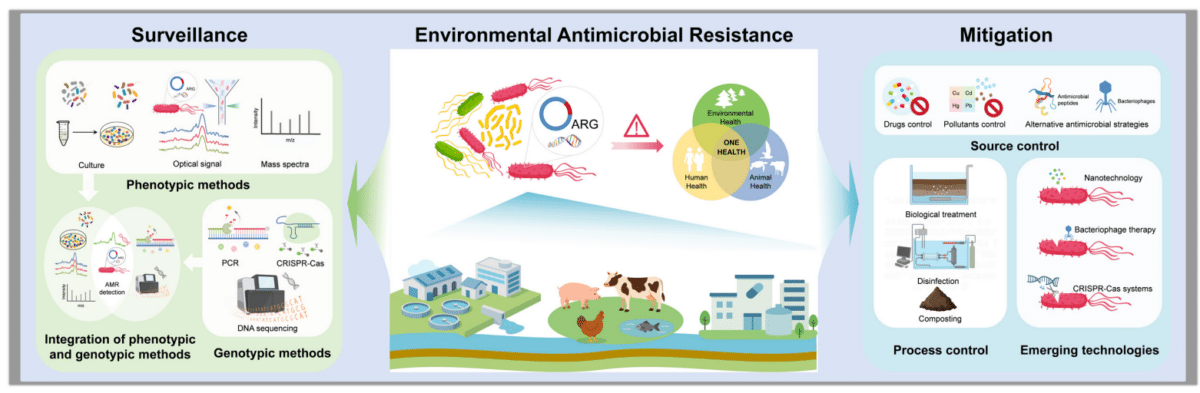
Huilin Zhang, Yuqiu Luo, Xinyu Zhu & Feng Ju)
Stopping Superbugs Of The Future
Researchers who study antibiotic resistance identified three factors that determine how dangerous resistance genes are: whether they can jump between bacteria easily, whether they’re carried by bacteria that cause human disease, and whether they confer resistance to multiple drug types.
Pharmaceutical factory waste scores high on all three. The resistance genes are mobile, they’re found in disease-causing bacteria, and they often provide resistance to several antibiotics at once.
Right now, pharmaceutical manufacturing operates with less oversight for environmental resistance than hospitals or farms. Changing that would require mandatory monitoring of resistance genes in factory wastewater, treatment standards that address living contamination (not just chemicals), and transparency about what’s being released and where.
It would also require international cooperation, because antibiotic manufacturing is a global supply chain. A factory in one country can contribute to resistance problems in distant healthcare systems.
The technology to clean up pharmaceutical pollution exists. What’s missing is the regulatory framework to require it. Until that changes, antibiotic factories will keep functioning as superbug factories, and the resistance crisis will keep getting worse.
Paper Notes
Limitations
The reviewed research relies primarily on case studies from specific geographic regions, particularly China and other areas with concentrated pharmaceutical manufacturing. Resistance patterns from pharmaceutical facilities in other regions may differ. The studies measure resistance genes and bacteria in environmental samples but don’t always directly track how this pollution contributes to human infections, making the clinical relevance sometimes indirect. Standardized monitoring protocols for pharmaceutical waste are lacking globally, making direct comparisons across facilities and regions difficult.
Funding and Disclosures
This review article was supported by the Muyuan Laboratory under Grant No. 1136022401, Zhejiang Provincial Natural Science Foundation of China under Grant No. LR22D010001, and National Natural Science Foundation of China under Grant No. 51908467. The authors declared no conflicts of interest.
Publication Details
Authors: Huilin Zhang, Yuqiu Luo, Xinyu Zhu, and Feng | Journal: Biocontaminant, Volume 1, 2025 | Paper Title: “Environmental antimicrobial resistance: key reservoirs, surveillance and mitigation under One Health” | DOI: https://doi.org/10.48130/biocontam-0025-0023 | Affiliations: Environmental Microbiome and Biotechnology Laboratory, Center of Synthetic Biology and Integrated Bioengineering, School of Engineering, Westlake University, Hangzhou, China; Zhejiang Key Laboratory of Low-Carbon Intelligent Synthetic Biology, Research Center for Industries of the Future, Westlake University; Institute of Advanced Technology, Westlake Institute for Advanced Study; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, School of Life Sciences, Westlake University







