(Credit: Kitsawet Saethao/Shutterstock)
In A Nutshell
- Scientists discovered a 5,500-year-old form of the bacteria that causes syphilis in remains from Colombia, the oldest evidence of these diseases by more than 3,000 years.
- The ancient strain represents an entirely new branch that split off roughly 13,700 years ago, before modern syphilis, yaws, and bejel bacteria evolved into separate forms.
- The infected individual showed no bone damage, proving these diseases can remain hidden in ancient remains and suggesting many more cases went undetected throughout history.
- The finding confirms these infections were already spreading in the Americas thousands of years before European contact, adding crucial evidence to the centuries-old debate about syphilis origins.
A hunter-gatherer who died in what is now Colombia more than 5,000 years ago carried a secret in their bones. Scientists analyzing the remains recently discovered DNA from an ancient form of the bacteria that causes syphilis and related diseases. The finding pushes back the timeline of when and where these infections emerged.
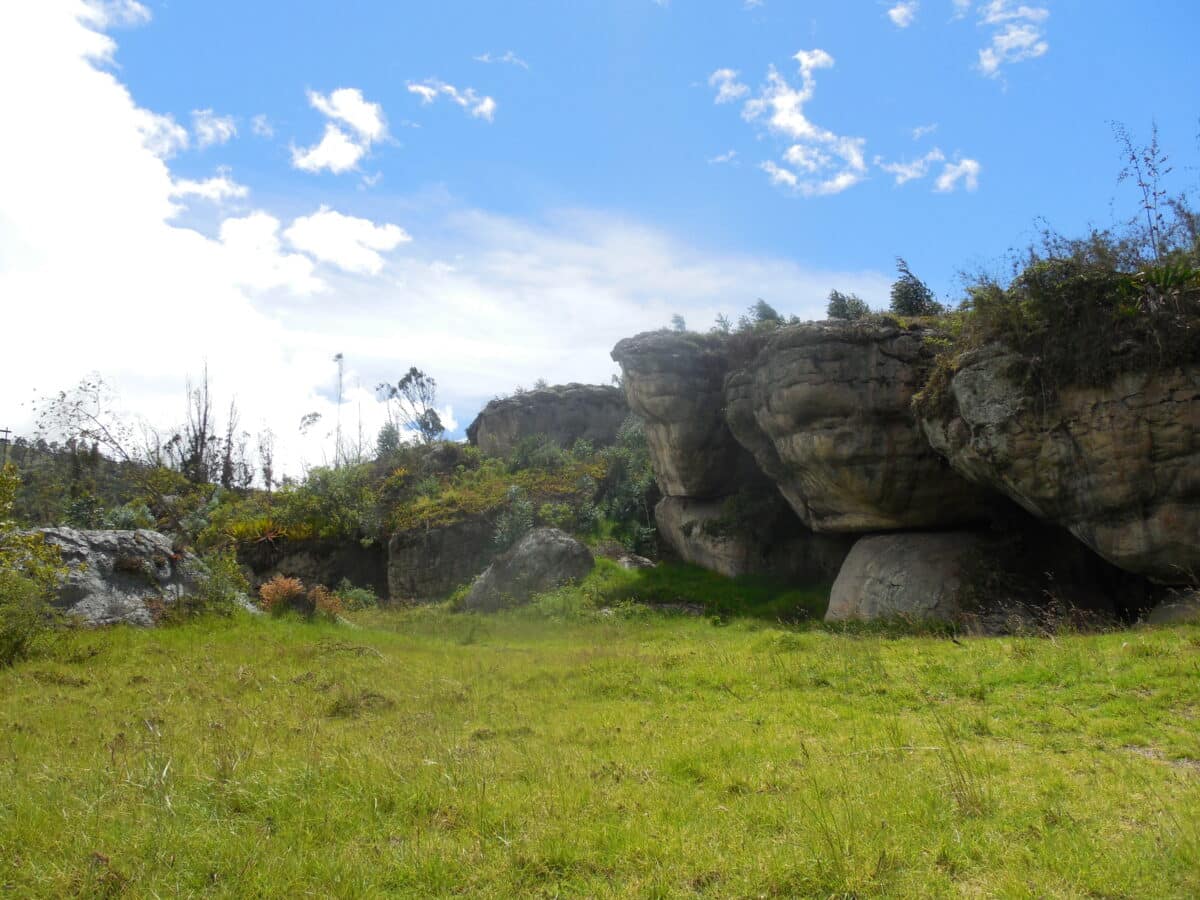
The individual, a middle-aged man buried at the Tequendama I rock shelter in the highlands near Bogotá around 3500 BCE, lived during a time when small groups of people moved across the landscape hunting deer and armadillo, gathering wild plants, and slowly experimenting with early forms of agriculture. He showed no signs of the bone damage typically caused by these infections, yet his remains harbored enough bacterial DNA for researchers to reconstruct most of the pathogen’s genetic blueprint.
The research team was surprised to find an entirely new form of Treponema pallidum that scientists estimate split from all known disease-causing strains roughly 13,700 years ago. That’s around the same time that the first humans were spreading across the Americas.
The Accidental Discovery
Initially, the research team wasn’t looking for ancient diseases. They were sequencing DNA to study human migration patterns when their screening software flagged something unexpected: genetic material from Treponema pallidum, the corkscrew-shaped bacterium responsible for syphilis (sexually transmitted), yaws (spread through skin contact), and bejel (transmitted through shared utensils or kissing).
The bacterial DNA represented just 0.0019% of everything in the sample, a tiny fraction buried among fragments of human DNA, soil bacteria, and environmental contamination. Deep sequencing of 1.5 billion DNA fragments allowed the team to piece together about 79% of the ancient pathogen’s genetic blueprint, enough to understand where it fits in the bacterial family tree.
This aspect of the discovery confounded study authors. Infections from these bacteria typically only damage bones in later stages when bacterial counts are already low. Most infected people never develop skeletal changes at all. Finding the pathogen in someone with no visible bone damage suggests either an early infection or a form of the disease that primarily affected skin and soft tissues.
A Missing Evolutionary Branch
When researchers compared the ancient strain (which they named TE1-3) to 106 modern and historical bacterial genomes, they found something unprecedented. TE1-3 sits on its own branch of the family tree, having split off before the ancestors of modern syphilis, yaws, and bejel diverged from each other.
In other words, it represents a lineage that was already ancient when the bacteria we know today were just beginning to evolve into distinct forms.
The ancient genome contains genes associated with disease-causing ability in modern strains, suggesting it could cause similar illness. But it also has two unique genetic changes affecting how the bacteria sense and respond to their environment. The effects of these changes remain unknown without laboratory testing.
Scientists can’t yet identify genetic markers that distinguish sexually transmitted syphilis from skin-contact infections like yaws. The bacteria are over 99% identical genetically despite causing different disease patterns. This means TE1-3 could represent an ancestor of syphilis, yaws, bejel, or even pinta (a skin-only infection for which no genetic data exists).
Ancient Infections in the Americas
The discovery, published in Science, pushes back the known timeline of these infections in the Americas by more than 3,000 years. Previous oldest samples dated to about 2,000 years ago from Brazil and Mexico.
Dating the evolutionary split is tricky with ancient DNA, but the researchers estimate TE1-3’s lineage diverged around 13,700 years ago, give or take several thousand years. That timeframe coincides with humans spreading throughout the Americas after crossing from Siberia between 15,000 and 20,000 years ago.
Modern forms of the bacteria appear to have diverged from each other much more recently, around 6,000 years ago. This suggests the pathogen was evolving alongside human populations for thousands of years, potentially adapting to different climates, transmission routes, and host populations.
The Sabana de Bogotá region where this individual lived was home to mobile hunter-gatherer groups during the Middle Holocene period. They hunted everything from large deer to small mammals like rabbits and guinea pigs. Interestingly, some rabbit species carry a closely related bacterium called Treponema paraluisleporidarum. While that species doesn’t naturally infect humans, it raises questions about the ancient origins of these pathogens and whether they once jumped between species.

The Syphilis Origins Debate
For centuries, historians have argued about whether syphilis existed in Europe before 1492. The “Columbian hypothesis” holds that Columbus’s crew brought the disease back from the Americas, sparking the devastating epidemics that swept through Europe in the late 1500s. The “pre-Columbian hypothesis” argues the disease already existed in Europe but went unrecognized.
This discovery doesn’t settle that debate, but it confirms treponemal bacteria were circulating in the Americas thousands of years before European contact. Skeletal evidence from sites across South America shows these infections affected indigenous populations throughout the region during this time period.
What scientists still can’t determine is when sexually transmitted syphilis specifically emerged. The bacteria that cause syphilis, yaws, and bejel are so similar genetically that they’re classified as subspecies of the same organism, distinguished mainly by how they spread rather than their DNA.
Answering that question will require combining ancient DNA evidence with demographic patterns in skeletal collections. If pre-Columbian populations show infections primarily in children, that suggests non-sexual transmission. A shift toward more adult cases after European contact might signal the emergence or introduction of sexually transmitted forms.
Why This Matters
This single genome transforms scientists’ understanding of how long these bacteria have been evolving with human populations. Many bone collections originally studied for human genetics may harbor hidden pathogen DNA that could reveal even more about ancient disease patterns.
Each new ancient genome provides a calibration point for reconstructing when different infectious diseases emerged and how they adapted to human hosts. As DNA sequencing becomes cheaper and metagenomic screening more routine, researchers expect to uncover disease histories that left no visible trace on ancient skeletons.
The discovery also raises intriguing questions about disease transmission. These infections typically require large, dense populations to maintain chains of transmission. Yet this individual lived in a small, mobile community. Either the bacteria evolved ways to persist in small groups through long latency periods, or there was far more contact between communities than archaeologists previously recognized.
For a pathogen that has shaped human history, from devastating 16th-century epidemics to modern public health challenges, understanding its deep evolutionary roots matters. Each ancient genome is a window into how infectious diseases have traveled with humans, adapted to our bodies, and left their mark on our species for thousands of years.
Paper Summary
Limitations
The study faces several constraints. The ancient genome’s low coverage means researchers captured only a subset of genetic variants. Using modern reference genomes for comparison may have skewed results, making the ancient strain appear more similar to contemporary bacteria than it truly was. The team tested multiple approaches but acknowledged their divergence date estimates likely represent minimum bounds. Without closely related modern strains for comparison, completely eliminating these biases proves difficult.
Laboratory testing would be needed to determine the functional effects of unique mutations in TE1-3. Genetics alone cannot reveal whether this strain caused sexually transmitted syphilis or skin-contact infections. The study examines only a single ancient genome, making it impossible to assess how common this form was during the Middle Holocene or whether the lineage went extinct versus persisting in unsampled modern populations. The individual’s lack of skeletal pathology prevents determining infection stage or tissue distribution.
Funding and Disclosures
This research was supported by the National Science Foundation Doctoral Dissertation Improvement Grant (award 2141920) to N.B.; European Research Council (grant agreement 679330) and Swiss National Science Foundation (grant PP00P3_176977) to A.-S.M.; Fundación de Investigaciones Arqueológicas Nacionales, Colombia (award 2009-409) to M.D.; National Institute of General Medical Sciences of the National Institutes of Health (award R35GM142939) to C.E.G.A.; European Research Council StG (grant agreement 948800, PaleoMetAmerica), Institut Pasteur and CNRS UMR 2000 funding, and INCEPTION program Investissement d’Avenir (grant ANR-16-CONV-0005) to N.R.; and Horizon Marie Skłodowska-Curie (award 210883809, PATHOGEN) to E.A.N. The authors declare no competing interests.
Publication Details
The study was authored by Davide Bozzi, Nasreen Z. Broomandkhoshbacht, Miguel Delgado, Jane E. Buikstra, Carlos Eduardo G. Amorim, Kalina Kassadjikova, Melissa Pratt Estrada, Gilbert Greub, Nicolas Rascovan, David Šmajs, Lars Fehren-Schmitz, Anna-Sapfo Malaspinas, and Elizabeth A. Nelson. Author affiliations include University of Lausanne (Department of Computational Biology and Swiss Institute of Bioinformatics); University of California Santa Cruz (UCSC Paleogenomics, Department of Anthropology, and UCSC Genomics Institute); Universidad Nacional de La Plata and CONICET, Argentina; Fudan University (School of Life Sciences); Arizona State University (School of Human Evolution and Social Change); California State University Northridge (Department of Biology); Instituto Colombiano de Antropología e Historia (ICANH), Bogotá; University of Lausanne and University Hospital Center CHUV (Institute of Microbiology and Service of Infectious Diseases); Institut Pasteur (Microbial Paleogenomics Unit), Paris; Masaryk University, Czech Republic (Department of Biology); and Southern Methodist University (Department of Anthropology). The research was published in Science on January 22, 2026. DOI: 10.1126/science.adw3020.







