
(© DOC RABE Media - stock.adobe.com)
BOSTON — In a world where microscopic battles rage unseen, one cunning adversary stands out – the herpes simplex virus (HSV). This master of disguise, affecting billions worldwide, has long puzzled scientists with its uncanny ability to dodge our best medications. But now, thanks to groundbreaking research from Harvard Medical School, scientists are finally peeking behind the viral curtain.
HSV is a virus that is so adaptable, it’s like a chameleon on steroids. Known for causing those pesky cold sores, HSV can transform itself faster than you can say “antiviral.” This shape-shifting prowess isn’t just annoying – it’s potentially dangerous, especially for those with weakened immune systems.
Enter the scientific sleuths from Harvard, armed with a microscope so powerful it can freeze-frame molecules mid-dance. Using this high-tech camera, called cryogenic electron microscopy (cryo-EM), they’ve captured HSV’s secret weapon in action.
The viral vanishing act, it turns out, isn’t just about changing colors, according to the study published in the journal Cell. It’s an intricate ballet performed by tiny protein machines called polymerases. These molecular acrobats are responsible for the virus’s ability to copy itself and spread. And boy, are they flexible!
“Our findings show that we have to think beyond targeting the typical drug-binding sites,” says Jonathan Abraham, the study’s senior author and an associate professor at Harvard Medical School, in a media release. “This really helps us see drug resistance in a new light.”
Here’s where it gets interesting. Imagine trying to catch a robber, only to find out they’re not hiding where you expected. That’s exactly what these researchers discovered. The virus doesn’t just change at the spots where our drugs attack. Instead, it makes tweaks far away from these sites, altering how its copying machine moves and works.
This discovery flips the script on everything we thought we knew about viral drug resistance. It’s not just about changing locks – it’s about redesigning the whole door.
Don’t worry, this isn’t all doom and gloom. Understanding these viral gymnastics opens up a whole new playbook for fighting back. Instead of just aiming for the obvious targets, future treatments might try to freeze these proteins in their tracks, stopping the virus’s shape-shifting shenanigans before they start.
This breakthrough isn’t just about herpes. The insights gained here could be a game-changer in our ongoing battle against all sorts of viruses that like to play dress-up.
As we continue to unmask these microscopic masters of disguise, one thing is clear: in the world of viruses, it’s not just about what you are, but how you move. And now, thanks to some clever scientific detective work, we’re one step closer to catching these shifty characters in the act.
“I’ve worked on HSV polymerase and acyclovir resistance for 45 years. Back then I thought that resistance mutations would help us understand how the polymerase recognizes features of the natural molecules that the drugs mimic,” says study co-author Donald Coen, professor of biological chemistry and molecular pharmacology at HMS. “I’m delighted that this work shows that I was wrong and finally gives us at least one clear reason why HSV polymerase is selectively inhibited by the drug.”
Paper Summary
Methodology
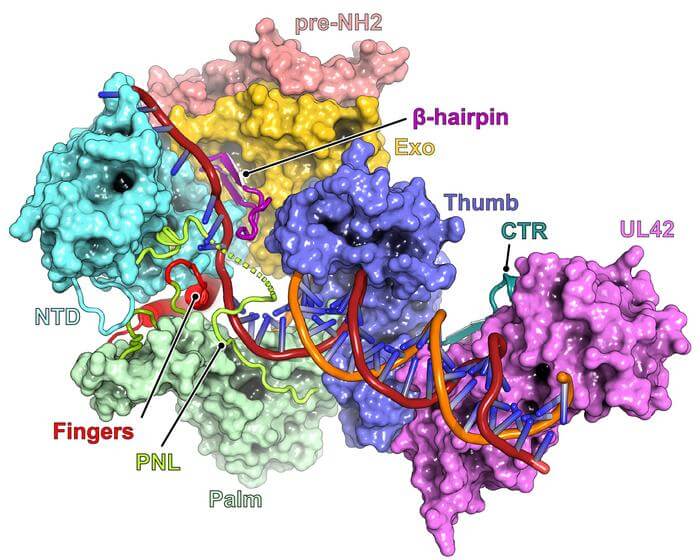
The researchers used a combination of cryo-EM and molecular dynamics simulations to visualize the DNA polymerase enzyme at an atomic level. Cryo-EM involves rapidly freezing the enzyme and then using an electron microscope to capture images of it in different states. These images were then used to create detailed 3D models of the enzyme, showing how it changes shape during the replication process and how it interacts with antiviral drugs.
Molecular dynamics simulations were employed to further explore how these mutations affect the enzyme’s behavior. These simulations provide a dynamic view of the enzyme, showing how it moves and fluctuates in response to the presence of DNA or drugs. By comparing the behavior of the wild-type (non-mutated) enzyme with that of the mutated versions, the researchers could pinpoint exactly how these mutations lead to drug resistance.
Key Results
The study’s results offer a new perspective on antiviral drug resistance. The detailed structures of the HSV DNA polymerase, both in its drug-bound and drug-free forms, reveal that mutations leading to resistance often do not directly block the drug from binding. Instead, they subtly alter the enzyme’s conformational dynamics, making it less likely for the drug to bind effectively.
For instance, the W781V mutation doesn’t prevent the drug from binding outright but increases the flexibility of the enzyme, making it more difficult for the drug to lock the enzyme into a non-functional state. This nuanced understanding of resistance mechanisms highlights the importance of targeting the dynamic nature of these enzymes in the design of future antiviral drugs.
Study Limitations
The research focused primarily on the HSV DNA polymerase, so the findings may not directly apply to other viruses. Additionally, the study mainly used in vitro (test tube) models, so further research is necessary to confirm these findings in living organisms. Moreover, while the study explains how these mutations confer resistance, it does not address how quickly these mutations can arise in a viral population, which is crucial for understanding the spread of resistance.
Discussion & Takeaways
The findings from this study have significant implications for the future of antiviral drug development. Understanding the conformational dynamics of DNA polymerase could lead to the creation of more effective drugs that can overcome resistance. For example, designing drugs that target the enzyme in both its open and closed states could help prevent the virus from evading treatment.
Moreover, this research underscores the importance of considering the dynamic nature of enzymes when developing drugs. Instead of just looking at how a drug binds to a static enzyme structure, scientists need to consider how the enzyme moves and changes shape. This approach could lead to the development of drugs that are more resilient to resistance, potentially extending their useful lifespan.
Funding & Disclosures
This study was supported by grants from the National Institutes of Health (NIH) and the Centers for Integrated Solutions in Infectious Diseases. The researchers declare no competing interests.







