ITHACA, N.Y. — Bad news, there are now even more forms of listeria which can contaminate food. Researchers from Cornell University say they’ve stumbled across five never-before-seen relatives of this potential deadly bacteria. The good news is the discovery is giving health officials more knowledge to help keep your food safe.
Study authors add their findings are particularly important because one of these listeria species is completely different from typical strains. While this family of bacteria usually moves a lot, the new species, L. immobilis, lacks the ability to move. Researchers say this means the tests for listeria contamination which focus on movement are about to get a serious re-write.
“This research increases the set of listeria species monitored in food production environments,” says lead author Catharine R. Carlin, a doctoral student in food science, in a university release. “Expanding the knowledge base to understand the diversity of listeria will save the commercial food world confusion and errors, as well as prevent contamination, explain false positives and thwart foodborne outbreaks.”
Until now, scientists believed motility was a common trait among strains related to L. monocytogenes, a well-known foodborne germ. Since various bacteria species can exist in environments that support L. monocytogenes, food production facilities have to monitor for all kinds of listeria.
Listeria monocytogenes can quickly spread through a factory, leading to outbreaks among the consumers eating this food. Listeriosis can kill between 20 and 30 percent of the people infected, even if they’re taking antibiotics. According to the CDC, about 1,600 people contract listeriosis each year and 260 die from it.
So what does this mean for your trip to the grocery store?
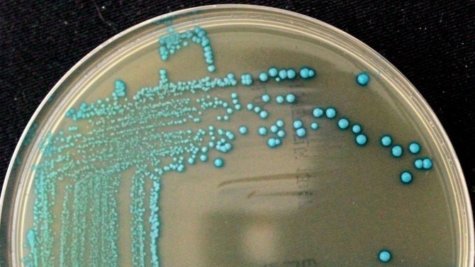
Researchers admit consumers probably won’t notice a difference as they browse their local produce aisles. However, the discovery of five new species of listeria means professional health inspectors can now keep more germs out of your food.
“This paper describes some unique characteristics of listeria species that are closely related to listeria monocytogenes, which will be important from an evolutionary perspective and from a practical standpoint for the food industry,” explains co-author Martin Wiedmann, Ph.D. ’97, the Gellert Family Professor in Food Safety and Food Science. “Likely, some tests will need to be re-evaluated.”
“This will help us to get better about identifying listeria monocytogenes, and not misidentifying it as something else,” Wiedmann adds.
What other species of bacteria are lurking near food?
Wiedmann’s team collected soil samples from across the United States and agricultural water samples from New York state during this study. Researchers uncovered 27 listeria microorganisms they could not classify and connect to any known bacteria species. Using whole-genome sequence-based tests, the team discovered the germs belong to five new clusters of listeria.
Along with the aptly-named L. immobilis (due to its inability to move), scientists named three of the species in honor of listeria researchers:
- L. cossartiae for Pascale Cossart, a bacteriologist at the Pasteur Institute of Paris
- L. farberi for Jeff Farber, a microbiologist at the University of Guelph in Canada
- L. portnoyii for Daniel Portnoy, a microbiologist at the University of California-Berkeley
Study authors named the final unique listeria species L. rustica; coming from the Latin word “rusticus” which honors the germ’s rural roots. Interestingly, Wiedmann’s group has unearthed 13 of the 26 listeria species discovered since 2010.
“When you’re inspecting the environments of food processing plants or restaurants, you need to know the pathogenic listeria from the non-pathogenic species,” Wiedmann concluded. “You need to tell the good guys from the bad guys.”
The study appears in the International Journal of Systematic and Evolutionary Microbiology.
